NexaMass-V3-Struct
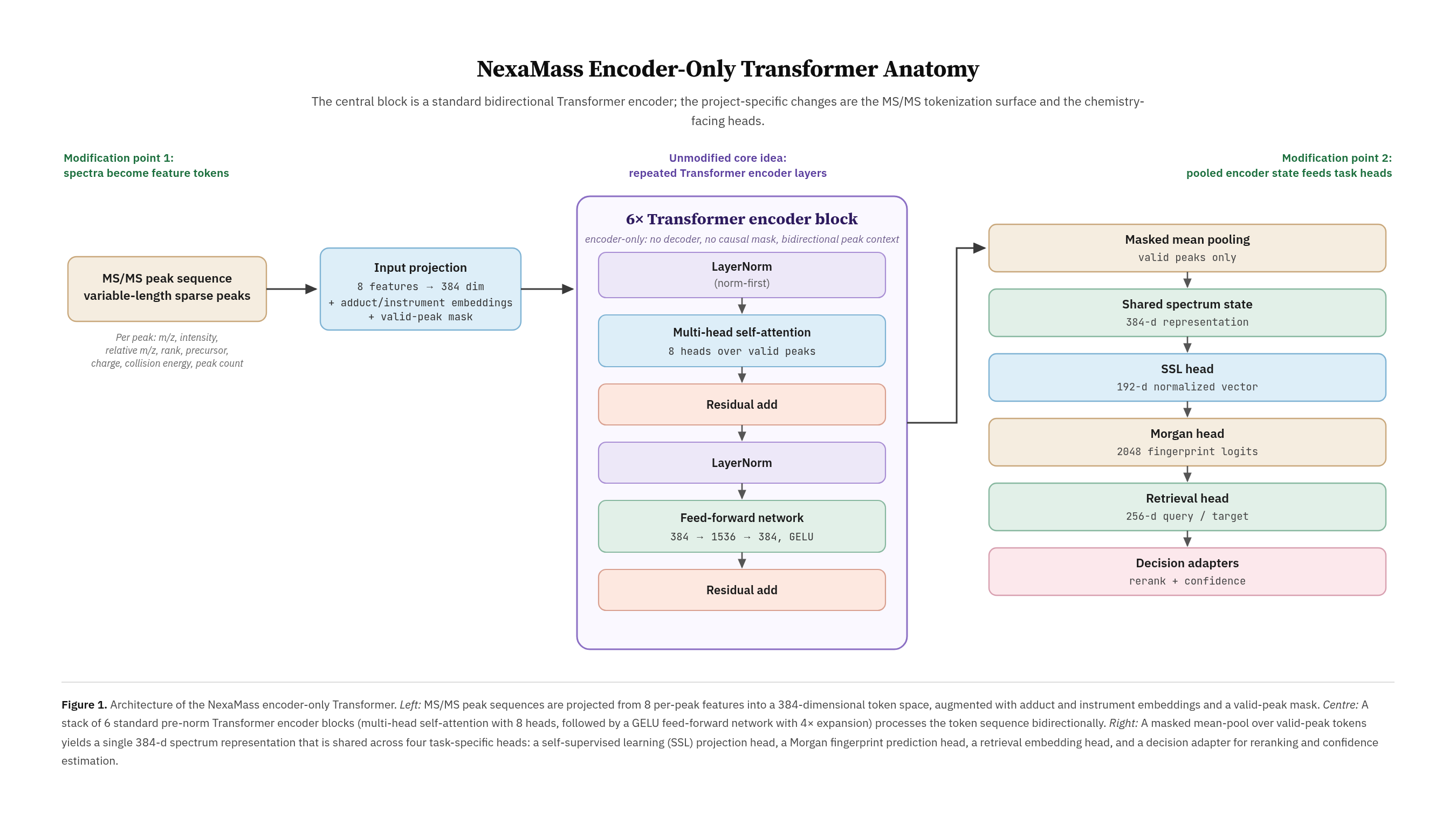
NexaMass-V3-Struct is a compact MS/MS spectral encoder for structure-aware representation learning and candidate-bank molecular inference. It maps tandem mass spectra into a learned spectral embedding and predicts RDKit Morgan fingerprint probabilities that can be compared against candidate molecular fingerprints. The model is intended for spectrum embedding, candidate narrowing, structure-aware retrieval research, and confidence/abstention experiments. It is not a de novo molecule generator and should not be used as a standalone top-1 molecular identifier.
The model was developed as part of the Nexa MS/MS pipeline. The foundation checkpoint, NexaMass-V3, was trained as a self-supervised spectral encoder over an approximately 201M-spectrum phase-1 campaign. The structure-aligned checkpoint, NexaMass-V3-Struct, adapts that encoder to a corrected labeled surface with real RDKit Morgan fingerprint targets. The main finding is that a small encoder can carry useful structure signal and support candidate-bank narrowing, while exact local ranking and calibrated confidence remain separate downstream problems.
Model Summary
NexaMass-V3-Struct is an encoder-only transformer with 14,106,690 trainable parameters. The full training checkpoint is approximately 153 MB in object storage, while the public model-state weights are approximately 54 MB per checkpoint and are provided in both PyTorch and Safetensors formats, making the released model substantially smaller than many public MS/MS representation or chemistry foundation checkpoints. The model uses 256 peak tokens, an input feature width of 8, hidden dimension 384, 6 transformer layers, 8 attention heads, feed-forward dimension 1536, dropout 0.1, SSL projection dimension 192, RDKit Morgan fingerprint dimension 2048, target projection dimension 256, and retrieval MLP hidden dimension 512.
The model contains a spectral transformer backbone, an SSL projection head, a Morgan fingerprint structure head, a spectrum-side retrieval query projection, a fingerprint-side target projection, and experimental retrieval/reranking heads. The strongest current inference surface is the Morgan fingerprint prediction and candidate-bank decode path. The trained retrieval projection is included for research, but it is not promoted as a reliable final decision layer.
Intended Use
Use this model to embed MS/MS spectra, build nearest-neighbor or clustering analyses, predict Morgan fingerprint probability vectors from spectra, score spectra against candidate banks, inspect structure-family neighborhoods, and develop ranking or confidence adapters over a frozen spectral encoder. The model is also suitable for research into structure-aware MS/MS representation learning and candidate narrowing.
Do not use this model as an unrestricted de novo structure generator. Do not treat the top-ranked candidate as a production-grade molecular identification without external validation. Do not treat the raw confidence or score gap as calibrated probability. Candidate outputs should be validated with chemistry-aware checks such as RDKit sanitization, formula and exact-mass agreement, precursor/adduct compatibility, fingerprint similarity, and spectrum-to-candidate consistency.
The preferred public inference files are weights/NexaMass-V3-Struct-model_state.safetensors and weights/Final_V3-model_state.safetensors. PyTorch model-state .pt fallbacks are included for compatibility. Optimizer-bearing full training checkpoints are retained in Wasabi/object storage and are not part of the public Hugging Face payload.
Inference Contract
The recommended inference path is spectrum-to-embedding plus candidate-bank decoding. A spectrum is converted into a normalized embedding and a 2048-dimensional Morgan fingerprint probability vector. That vector is compared against candidate molecular fingerprints from a known bank. The output should be a ranked shortlist with validation metadata, not a single unqualified structure claim.
A useful production-facing record should include the predicted candidate structure, canonical SMILES, formula, exact mass, precursor mass error, adduct compatibility, RDKit validity, fingerprint similarity, rank, confidence band, ambiguity band, and optional fragment-support notes. This framing is intentionally conservative: the model narrows candidate hypotheses; it does not generate molecules from unconstrained chemical space.
Training Data
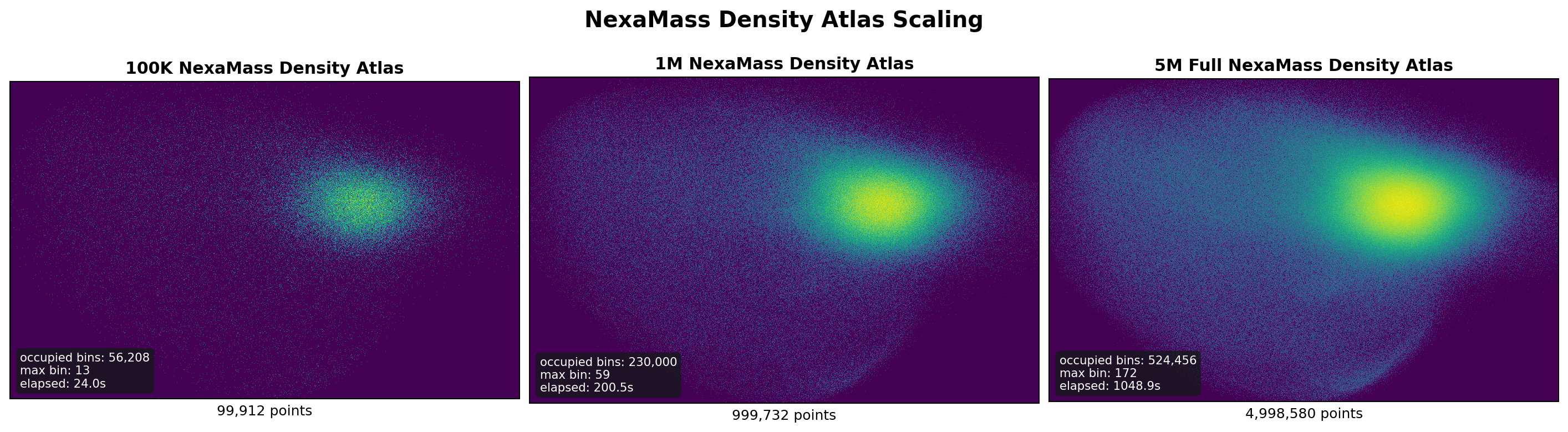
The foundation encoder was derived from the public GeMS MS/MS dataset released by the DreaMS authors at roman-bushuiev/GeMS on Hugging Face: https://huggingface.co/datasets/roman-bushuiev/GeMS. The GeMS data were processed into deterministic Parquet shard windows for the NexaMass training pipeline. The phase-1 pretraining campaign processed approximately 201M spectra across three scale windows. The training stack used self-supervised spectral objectives with light evaluation during training and deeper holdout evaluation at phase boundaries.
The molecular structure alignment stage used a corrected labeled surface where RDKit Morgan targets were present and enforced. Broad foundation pretraining metrics should be interpreted as SSL representation evidence. Molecular grounding should be interpreted through the corrected structure-alignment run rather than through unlabeled foundation shards.
Training Procedure
The foundation model was trained as an encoder-only spectral transformer using a self-supervised contrastive objective. The promoted foundation checkpoint, NexaMass-V3, reached stable representation behavior over the phase-1 campaign, with contrastive loss improving from 0.6206 to 0.3373 across V1-V3 and embedding standard deviation remaining stable near 0.072.
The structure-aligned checkpoint, NexaMass-V3-Struct, resumed from the foundation encoder and trained against real RDKit Morgan fingerprint targets. The training objective combined a small SSL preservation term with structure prediction and retrieval-facing losses. This stage validated that the foundation encoder could attach to molecular structure targets without representation collapse.
Evaluation
The structure-aligned checkpoint reached validation structure BCE 0.0653, validation fingerprint cosine 0.4255, and embedding standard deviation 0.0722 in the V26 alignment run. A candidate-bank structure gallery over 20 displayed validation examples matched the ground-truth candidate identity in 11 cases through the fingerprint decode path. The same gallery matched 0 exact identities through the trained retrieval projection, which is why the card treats ranking as an open decision-layer problem rather than a solved property.
On a MassSpecGym retrieval adapter using the frozen V3 projected-dot scorer, the model reached validation Hit@1 0.1162, Hit@5 0.1915, and Hit@20 0.3328. On the MassSpecGym test dataloader under Hit@k-only evaluation, it reached Hit@1 0.0627, Hit@5 0.1753, and Hit@20 0.3505. This Hit@20-only adapter comparison places the model above lower MassSpecGym retrieval baselines such as Random, DeepSets, Fingerprint FFN, and DeepSets with Fourier features, but below specialized retrieval systems such as MIST. The result should be interpreted as evidence of useful top-k narrowing, not leaderboard-grade retrieval.
Internal larger-bank pressure tests also support a bounded interpretation. On a roughly 29k unique-identity candidate bank, the frozen projected-dot scorer beat random and compressed candidate rank substantially, but exact local discrimination remained weak. The best current read is that the model contains real structure and retrieval signal, while top-1 ranking and calibrated confidence require additional downstream machinery.
Limitations
NexaMass-V3-Struct has several important limitations. The labeled structure-alignment surface is smaller than the full self-supervised pretraining corpus, so broader label-rich evaluation is needed before claiming chemical-space-wide generalization. The current trained retrieval projection does not reliably convert structure signal into exact local ordering. Confidence is not calibrated and should be used only as a research signal unless paired with abstention curves and ambiguity-tiered validation. The model does not perform unrestricted de novo generation and does not directly decode SMILES or molecular graphs.
The safest use is candidate narrowing with explicit uncertainty. The model can help reduce a large candidate space to a smaller set of chemically plausible hypotheses, but final interpretation should remain analyst-reviewed and chemistry-validated.
Ethical And Practical Considerations
MS/MS structure inference can affect downstream scientific interpretation. Users should validate candidate outputs against experimental context, known chemistry, instrument settings, precursor information, and independent evidence. This model is intended as a research and decision-support tool, not as an autonomous chemical identification authority.
Citation
If you use this model, cite the NexaMass project release and the accompanying technical report when available. Relevant background work includes DreaMS for self-supervised MS/MS representation learning, MassSpecGym for benchmark framing, CSI:FingerID for fingerprint-mediated candidate search, and related spectra-structure retrieval and de novo generation systems such as MIST, MSNovelist, CMSSP, CSU-MS2, MSBERT, Spec2Mol, and MS2Mol.
MassSpecGym Adapter
A safetensors-compatible MassSpecGym retrieval adapter is included under evaluation/massspecgym/. It loads weights/NexaMass-V3-Struct-model_state.safetensors, converts MassSpecGym tokenized spectra into the NexaMass batch contract, and reports Hit@k retrieval metrics through MassSpecGym's evaluator. The archived reference run reached test Hit@20 0.3505 with the frozen projected-dot scorer. This should be read as evidence of transferable top-k signal, not solved molecular ranking or calibrated confidence.
- Downloads last month
- 67
Datasets used to train AethronPhantom/NexaMass-V3-Struct
Evaluation results
- Hit@20 on MassSpecGymNexaMass External Benchmark Adapter (Frozen V3)0.350